Health Informatics
- Biomedical Informatics, Biostatistics, and Data Science Resources and Services
- Center for Computational Biology and Bioinformatics
- Biostatistics, Epidemiology and Research Design Center
Velos eResearch is an integrated software system for managing clinical trials. The software links to the UCSD Health's Epic Electronic Medical Record System to provide improved information and integration for clinical research projects.
One module within this platform, called eSample, will track biological samples and link them to the Electronic Health Record.
A robust support team assists investigators in implementing their protocols, study budgets, and calendars.
Velos eResearch is a web-based system that supports:
It is mandatory that all new users attend Velos CRB training session, upcoming training schedule is available on the Velos Wiki (requires UCSD VPN Access). To reserve a space for the Training, please e-mail ctri-velos@ucsd.edu. Once users have attended a training session and submitted a completed and signed access request form, access to the Velos Application will be granted.
To request access for Velos Application follow the request services button on top of the page.
Select "Biomedical Informatics" / "Velos Access"
The Application Support Team conducts detailed in-depth hands-on Velos CRB training sessions twice a month.
Once granted Velos access, the Velos CRB training manuals are accessible within the Velos application under the question mark (?) icon in the upper right corner.
Training material (manuals and videos) are also available on the Velos Wiki (requires UCSD VPN Access) webpage.
Custom trainings tailored to your study needs can be provided: eSampling, budgeting, reporting etc.
For any questions regarding Velos (general inquiries, custom trainings etc), please contact the Velos Application Support Team (ctri-velos@ucsd.edu) or phone (858) 534-0555.

REDCap (Research Electronic Data Capture) is a secure, web-based application for building and managing online surveys and databases.
REDCap provides automated export procedures for seamless data downloads to Excel and common statistical packages (SPSS, SAS, Stata, R), as well as a built-in project calendar, a scheduling module, ad hoc reporting tools, and advanced features, such as branching logic, file uploading, and calculated fields.
While REDCap may be used to store PHI, it is not CFR 21 Part 11 compliant. If your project requires compliance with CFR 21 Part 11, you will need to use the Velos system.
REDCap is available to all UC San Diego faculty and staff, and to users outside the organization who have sponsorship from UC San Diego faculty. To gain access, all REDCap users must have UC San Diego Active Directory (AD) system credentials (contact ITS Service Desk for credentials)
To request access to REDCap, follow the request services button on top of the page.
Select "Biomedical Informatics"/ "REDCap Access"
Once granted access to the REDCap application, all users have access to extensive training videos, located on the REDCap home page.
The REDCap Wiki is a tool available to all REDCap users to post questions to the UCSD REDCap community.
For any other questions regarding REDCap, please contact the REDCap Application Support Team (ctri-redcap@ucsd.edu) or phone (858) 534-0555.

ENACT is a network of sites from the National Clinical and Translation Science Award (CTSA) Consortium. ENACT network allows researchers to explore and validate feasibility for clinical studies across CTSA sites. ENACT network has been created to harmonize EHR of each site linked by the Shared Health Research Information Network (SHRINE). ENACT provides investigators the ability to design and obtain aggregated counts of patients to identify eligible participants under specific inclusion and exclusion criteria. The real-time query of ENACT network to diverse CTSA sites across the United States significantly increases efficiency of clinical studies.
The ENACT network currently connects 39 participated CTSA sites, including all 5 UC medical centers, and more than 100 million patients. More sites are joining and at staging status.
For more information visit the ENACT Network information site.
To request access to the ENACT Network, please complete the BIDS Service Request. Select the “Access to Accrual to Clinical Trials (ACT/ENACT)” option.
The user-friendly ENACT SHRINE query interface is shown as the following. It simply consists of 4 panels. The Terms panel allows investigators to search the information of Demographics, Diagnosis, Laboratory tests, Medications, Procedures, and Visit details. The defined inclusion/exclusion criteria can be dragged and dropped to numerous groups on the Query Tool panel. Each group enables independent criteria date range and times of occurrence. The current and previous query results are shown in the lower panels. In addition, ENACT allows aggregate count breakdown for patient age, race, gender, and vital status, and provides graphic reports.
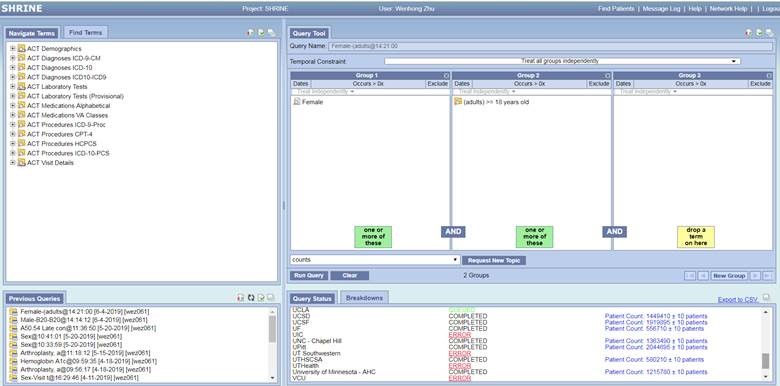
For any other questions regarding the ENACT Network, please contact the Application Support Team (ctri-support@ucsd.edu) or phone (858) 534-0555.

The Virtual Secure Workstation (VSW) offers researchers a secure way to store and access their research data. It allows the hosting of a desktop operating system on a centralized HIPAA-compliant server that is managed by the BIDS Unit. VSW access is limited to users with a valid UCSD Active Directory account and multi-factor authentication through Duo multi-factor authentication service. For extra security, the VSW environment is self-contained and allows no access to printers, internet, email or external drives.
VSW can also be customized to include your own licenses and/or applications.
Currently VSW is a free service provided by the BIDS Unit. To request VSW access please submit a BIDS Request and select "Virtual Secure Workstation (VSW)" Registration to the UC San Diego Health Duo multi-factor authentication service is also needed in order to log in to your VSW.
Data in the VSW shared folder is removed after 30 days – this shared folder is used to deliver results (such as DECS) to the end user and is not intended for long-term storage; users are expected to move the data to the VSW personal folder. Data in the VSW personal folder is preserved for the lifetime of the VRD account.
Data Import/Export
For all data migration requests, please email the Application Support Team (ctri-support@ucsd.edu) with the “VSW:” in the subject.
For import please include:
For export please include:
Data exports will only be allowed onto UCSD Health System controlled computers which are secure and encrypted.
For any other questions regarding the VSW, please contact the Application Support Team (ctri-support@ucsd.edu) or phone (858) 534-0555.

After much extended collaborative efforts between the ACTRI and MCC along with multiple research subject matter experts from other UCSD departments, we are now launching our new eRegulatory (eReg) services powered by the Florence application platform. The goal of implementing the Florence platform at UCSD is to enable us to move away from reliance on paper forms and physical research binders, thereby enhancing health research compliance through improving both our electronic regulatory and clinical documentation workflows.
New System Highlights
Service Fee Schedule - Effective 7/1/26
If you are interested in using UCSD’s Florence eReg platform, please send an email to actri-florence@health.ucsd.edu and we will reach out to you to initiate a consultation and discussion.
For more information on UCSD’s eReg Florence platform, you can visit our Florence Pulse website (requires AD Login): https://pulse.ucsd.edu/departments/clinical-research/Pages/Florence-eReg-System.aspx
All UCSD Research units are welcome to utilize Florence eRegulatory platform and can request study and user creations using the Florence Request Form. ACTRI BIDS Team manages this system.
Access the Florence eRegulatory platform SOP here.

The UCSD Biosignal Repository is a centralized, continuously maintained, multi-modal biosignal registry and research data infrastructure housed at the UC San Diego Clinical and Translational Research Institute (ACTRI). Led by Mattheus Ramsis, MD, Director of Cardiology Informatics, the repository captures and curates all forms of physiologic signal data generated within the UCSD health system, including electrocardiography, photoplethysmography, bedside waveforms, sleep study recordings, cardiopulmonary exercise data, pulmonary function signals, continuous glucose monitoring, vocal acoustic biomarkers, imaging data, and wearable device data, linked to rich clinical metadata from the Epic EMR.
The repository operates through three complementary data streams: collection of all available biosignal data from existing UCSD patient records; prospective ongoing capture of biosignals generated during routine clinical care, with no research-specific procedures added; and a governed secondary analysis access framework enabling qualified investigators to request de-identified data extracts for independent research. All data are de-identified prior to storage using HIPAA Safe Harbor standards and housed within a Trusted Research Environment meeting NIST SP 800-53 and HIPAA Security Rule requirements.
The repository is designed to serve as the foundational data infrastructure for translational research at UCSD, enabling discovery, predictive model development, multi-modal signal analysis, and equity-focused investigations across a health-system-scale patient population. Collaboration is actively invited from investigators across clinical departments, engineering, and data science.
For more information on Biosignal data and collaboration opportunities for research, please see the following contacts:
Dr. Matheus Ramsis
Founding Director Biosignal Repository
mramsis@health.ucsd.edu
Dr. Michael Hogarth
Director of ACTRI Biomedical Informatics Data Science
mihogarth@health.ucsd.edu
The Data Extraction Concierge Service (DECS) is a service that pulls data from the UCSD EPIC Electronic Medical Records system to provide UCSD patient and health data extracts for research purposes. This service allows to identifies specific patients, and extracts identified, de-identified, or limited patient-level datasets from electronic medical for clinical research projects. An approval by the Institutional Review Board (IRB) is required for extracting identified and limited datasets. Our support team will execute queries on the Clinical Data Warehouse for Research (CDWR) and return results to users.
Functionality
The data extract can include many different types of information, such as:
Any data visible in Epic can be included in an extraction request. Requests can be:
DECS delivers results to the requester through a dedicated Virtual Secure Workstation (VSW). By default, requested data are provided in a Microsoft Excel workbook (.xlsx), but other data formats are supported.
Access
To request DECS services follow the request services button on top of the page.
Select "Biomedical Informatics" / "Data Extraction Concierge Services"
The following are needed:
The details of your request:
Access to the DECS results VSW are only initially granted to the requester and/or PI. Only individuals listed in the IRB submission can be allowed access to the VSW. All VSW users require UCSD AD accounts.
Non-UCSD requests for DECS services normally require a UCSD P.I. sponsor, who will be responsible for the data generated. In addition, a legal data access agreement may be required.
Cost
The current recharge rate for DECS services is available here.
Generally, the final cost of a DECS request is proportional to its complexity: Number of elements (variables) to retrieve, number of inclusion and exclusion criteria, and how easy it is to retrieve the requested data.
As part of the initial review meeting, your DECS analyst can generally provide an estimate for the time and cost. It’s difficult to provide a firm quote, as each DECS request is unique – as work proceeds we can generally improve the estimate.
Some broad guidelines:
If your data budget is very tight, we can often organize your request into multiple “phases”, to meet your key requirements first, and the rest if the budget allows.
Contact and Help
For any other questions regarding DECS, please contact the Application Support Team (ctri-support@ucsd.edu) or phone (858) 534-0555.

The UC Health Data Warehouse (UCHDW) pulls a subset of the data from the Electronic Health Record system and allows investigators to query patient information in a HIPAA-compliant manner.
The UCHDW currently holds data on nearly 6 million patients seen at a UC facility since 2012. These patients received care from nearly 100,000 health care providers in over 200 million encounters, with nearly 200 million procedures, more than half a billion medication orders, and with over 2 billion vital signs measurements and test results. Over 600,000 of these patients are primary care patients.
Access to this information requires technical familiarity and ability to write code in R, Python, or Spark SQL, that queries the deidentified clinical data. The BMI team may assist to provision access directly to the UCHDW data. If assistance is needed in preparing the customized queries, there will be a recharge fee.
To request access, please sign up here and send questions to the Application Support Team (ctri-support@ucsd.edu).
Contact the IT Support Team at:
Weekly Office Hours - Data Extraction Concierge Service (DECS)
Thursdays 1:00PM - 2:00PM
Zoom link: https://uchealth.zoom.us/j/81732217241?pwd=Yk9rc0NPTGRTMnFxQkxhSTFaVEo2UT09&from=addon

Freezerworks is a powerful sample tracking, biorepository management, and freezer inventory program—available to you as a rechargeable service. Whether you're managing a clinical trial or a research study, Freezerworks helps streamline your data while maintaining regulatory compliance and data integrity.
What can you do with Freezerworks?
Track sample lineage for cell lines, tissues, blood products, and more
Configure secure access to data and freezers
Customize patient, sample, and aliquot records
Easily add fields, adjust screens and labels—no coding needed
View field interactions, calculations, dependencies, and smart lists
Import, modify, and search data across all fields
Manage workflows: check-in/out, requisitions, shipping, pooling, and testing
Maintain compliance with audit trails and multi-group support
Access via Desktop, REST API, or Browser Client
Reliable, affordable LIMS for clinical and research use
For more information about Biobanking services and FreezerWorks data integration, please contact the TRT division (ctri-lab@ucsd.edu) for consultation.
Phone: (858) 534-0555
ctri-velos@ucsd.edu
ctri-redcap@ucsd.edu
ctri-support@ucsd.edu
Please put application/service name in the e-mail subject.


Nguyen Trieu, MHIM
Chief Technology Officer
Phone: (858) 822-0111
